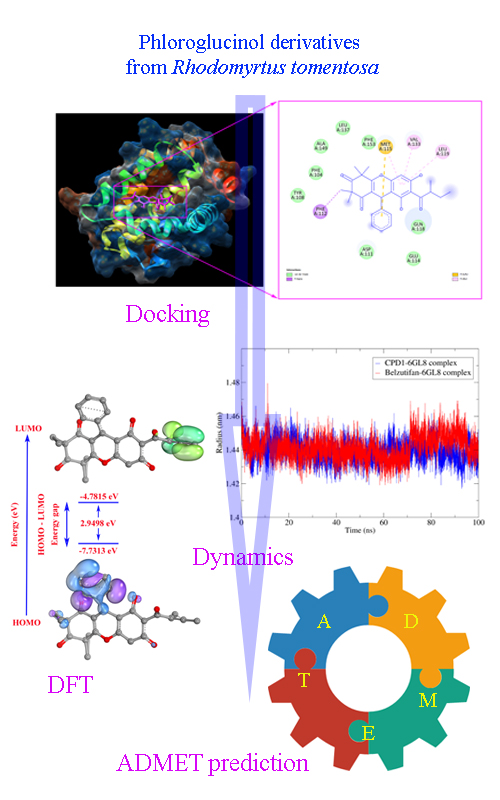
In silico modeling of selected phloroglucinol derivatives from Rhodomyrtus tomentosa: Mechanistic insights into their potential against cervical cancer
Main Article Content
Abstract
Cervical cancer remains a major cause of morbidity and mortality worldwide, driven primarily by persistent infection with high-risk HPV types. Natural phloroglucinol derivatives from Rhodomyrtus tomentosa have demonstrated anti-cervical cancer potential; however, their apoptosis-related mechanisms remain insufficiently characterized. In this work, selected phloroglucinol derivatives were evaluated as potential inhibitors of the anti-apoptotic Bcl-2 protein (PDB: 6GL8) through an integrated in silico pipeline that combines molecular docking, 100 ns molecular dynamics simulations, MM/GBSA binding free-energy estimation, ADMET profiling, and DFT-based electronic analysis. Docking prioritized CPD1 as the most favorable ligand (-37.36 kJ/mol), outperforming Belzutifan (-25.73 kJ/mol) and engaging the conserved binding pocket. MD trajectories supported stable complex formation across 100 ns. MM/GBSA further indicated stronger binding for CPD1 (ΔTOTAL = -138.78 ± 15.4 kJ/mol) relative to Belzutifan (-63.72 ± 14.31 kJ/mol), primarily due to more favorable gas-phase interactions, while maintaining a comparable solvation term. ADMET predictions suggested similar solubility but higher intestinal absorption for CPD1, alongside a hERG II alert that warrants cardiotoxicity-oriented optimization. DFT descriptors were consistent with the enhanced electrophilic character of CPD1. Collectively, CPD1 is computationally prioritized as a candidate scaffold for follow-up experimental validation and structure-guided refinement.
Downloads
Metrics
Article Details

This work is licensed under a Creative Commons Attribution 4.0 International License.

Authors retain copyright and grant the journal right of first publication with the work simultaneously licensed under a Creative Commons Attribution license 4.0 that allows others to share the work with an acknowledgement of the work's authorship and initial publication in this journal.
References
F. Bray, M. Laversanne, H. Sung, J. Ferlay, R. L. Siegel, I. Soerjomataram, A. Jemal, CA. Cancer J. Clin. 74 (2024) 229 (https://doi.org/10.3322/caac.21834)
K. S. Okunade, J. Obstet. Gynaecol. 40 (2020) 602 (https://doi.org/10.1080/01443615.2019.1634030)
M. Bida, B. Mosoane, B. Phakathi, M. Sebitloane, M. Z. Muallem, R. Hull, Z. Dlamini, in Cancer Sensitizing Agents for Chemotherapy, Z. Dlamini, Ed., Academic Press, Oxford, 2024, p. 17 (https://doi.org/10.1016/B978-0-443-28985-9.00005-7)
S. Ashique, M. Bhowmick, R. Pal, H. Khatoon, P. Kumar, H. Sharma, A. Garg, S. Kumar, U. Das, Adv. Cancer Biol. - Metastasis 10 (2024) 100114 (https://doi.org/10.1016/j.adcanc.2024.100114)
P. Ray, D. Paul, Pharmacol. Res. - Mod. Chinese Med. 12 (2024) 100497 (https://doi.org/10.1016/j.prmcm.2024.100497)
F. Qi, L. Zhao, A. Zhou, B. Zhang, A. Li, Z. Wang, J. Han, Biosci. Trends 9 (2015) 16 (https://doi.org/10.5582/bst.2015.01019)
Y. Zhou, J. Zheng, Y. Li, D. P. Xu, S. Li, Y. M. Chen, H. B. Li, Nutrients 8 (2016) 515 (https://doi.org/10.3390/nu8080515)
M. H. Kang, I. H. Kim, T. J. Nam, Int. J. Oncol. 45 (2014) 1036 (https://doi.org/10.3892/ijo.2014.2521)
Y. Peng, Y. Wang, C. Zhou, W. Mei, C. Zeng, Front. Oncol. 12 (2022) 819128 (https://doi.org/10.3389/fonc.2022.819128)
E. C. Aniogo, B. P. George, H. Abrahamse, Molecules 25 (2020) 5308 (https://doi.org/10.3390/molecules25225308)
N. V. Pervushin, G. S. Kopeina, B. Zhivotovsky, Biol. Direct 18 (2023) 69 (https://doi.org/10.1186/s13062-023-00431-4)
T. I. Adelusi, A. Q. K. Oyedele, I. D. Boyenle, A. T. Ogunlana, R. O. Adeyemi, C. D. Ukachi, M. O. Idris, O. T. Olaoba, I. O. Adedotun, O. E. Kolawole, Y. Xiaoxing, M. Abdul-Hammed, Informatics Med. Unlocked 29 (2022) 100880 (https://doi.org/10.1016/j.imu.2022.100880)
D. Jeong, W. S. Yang, Y. Yang, G. Nam, J. H. Kim, D. H. Yoon, H. J. Noh, S. Lee, T. W. Kim, G. H. Sung, J. Y. Cho, J. Ethnopharmacol. 146 (2013) 205 (https://doi.org/10.1016/j.jep.2012.12.034)
Y. B. Zhang, W. Li, L. Jiang, L. Yang, N. H. Chen, Z. N. Wu, Y. L. Li, G. C. Wang, Phytochemistry 153 (2018) 111 (https://doi.org/10.1016/j.phytochem.2018.05.018)
J. B. Murray, J. Davidson, I. Chen, B. Davis, P. Dokurno, C. J. Graham, R. Harris, A. Jordan, N. Matassova, C. Pedder, S. Ray, S. D. Roughley, J. Smith, C. Walmsley, Y. Wang, N. Whitehead, D. S. Williamson, P. Casara, T. Le Diguarher, J. Hickman, J. Stark, A. Kotschy, O. Geneste, R. E. Hubbard, ACS Omega 4 (2019) 8892 (https://doi.org/10.1021/acsomega.9b00611)
G. M. Morris, R. Huey, W. Lindstrom, M. F. Sanner, R. K. Belew, D. S. Goodsell, A. J. Olson, J. Comput. Chem. 30 (2009) 2785 (https://doi.org/10.1002/jcc.21256)
H. D. Nguyen, Not. Sci. Biol. 17 (2025) 12642 (https://doi.org/10.55779/nsb17312642)
D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark, H. J. C. Berendsen, J. Comput. Chem. 26 (2005) 1701 (https://doi.org/10.1002/jcc.20291)
N. Guex, M. C. Peitsch, Electrophoresis 18 (1997) 2714 (https://doi.org/10.1002/elps.1150181505)
V. Zoete, M. A. Cuendet, A. Grosdidier, O. Michielin, J. Comput. Chem. 32 (2011) 2359 (https://doi.org/10.1002/jcc.21816)
H. D. Nguyen, Indian J. Chem. 64 (2025) 383 (https://doi.org/10.56042/ijc.v64i4.15530)
M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente, E. Moreno, J. Chem. Theory Comput. 17 (2021) 6281 (https://doi.org/10.1021/acs.jctc.1c00645)
D. E. Pires, T. L. Blundell, D. B. Ascher, J. Med. Chem. 58 (2015) 4066 (https://doi.org/10.1021/acs.jmedchem.5b00104)
F. Neese, WIREs Comput. Mol. Sci. 15 (2025) e70019 (https://doi.org/10.1002/wcms.70019)
G. Knizia, J. E. Klein, Angew. Chemie Int. Ed. 54 (2015) 5518 (https://doi.org/10.1002/anie.201410637)
M. D. Hanwell, D. E. Curtis, D. C. Lonie, T. Vandermeersch, E. Zurek, G. R. Hutchison, J. Cheminform. 4 (2012) 17 (https://doi.org/10.1186/1758-2946-4-17)
J. Luo, Z. Q. Xue, W. M. Liu, J. L. Wu, Z. Q. Yang, J. Phys. Chem. A 110 (2006) 12005 (https://doi.org/10.1021/jp063669m)
R. Das, J. L. Vigneresse, P. K. Chattaraj, Int. J. Quantum Chem. 114 (2014) 1421 (https://doi.org/10.1002/qua.24706)
P. C. Agu, C. A. Afiukwa, O. U. Orji, E. M. Ezeh, I. H. Ofoke, C. O. Ogbu, E. I. Ugwuja, P. M. Aja, Sci. Rep. 13 (2023) 13398 (https://doi.org/10.1038/s41598-023-40160-2)
D. A. Schaller, C. D. Christ, J. D. Chodera, A. Volkamer, J. Chem. Inf. Model. 64 (2024) 8848 (https://doi.org/10.1021/acs.jcim.4c00905)
R. Patil, S. Das, A. Stanley, L. Yadav, A. Sudhakar, A. K. Varma, PLoS One 5 (2010) e12029 (https://doi.org/10.1371/journal.pone.0012029)
G. Bitencourt-Ferreira, M. Veit-Acosta, W. F. de Azevedo, in Docking Screens for Drug Discovery, W. F. de Azevedo Jr., Ed., Springer, New York, 2019, p. 79 (https://doi.org/10.1007/978-1-4939-9752-7_6)
H. Alonso, A. A. Bliznyuk, J. E. Gready, Med. Res. Rev. 26 (2006) 531 (https://doi.org/10.1002/med.20067)
L. Martínez, PLoS One 10 (2015) e0119264 (https://doi.org/10.1371/journal.pone.0119264)
M. L. Verteramo, M. M. Ignjatović, R. Kumar, S. Wernersson, V. Ekberg, J. Wallerstein, G. Carlström, V. Chadimová, H. Leffler, F. Zetterberg, D. T. Logan, U. Ryde, M. Akke, U. J. Nilsson, iScience 27 (2024) 109636 (https://doi.org/10.1016/j.isci.2024.109636)
H. D. Nguyen, Phys. Chem. Res. 13 (2025) 783 (https://doi.org/10.22036/pcr.2025.535946.2708)
H. Rohit, H. Tandon, J. Mol. Model. 31 (2025) 291 (https://doi.org/10.1007/s00894-025-06487-5)
B. Bashir, A. Z. Clayborne, Molecules 30 (2025) 2254 (https://doi.org/10.3390/molecules30112254).